What are FG beads® ?
Excellent tool for chemical biology
In pharmaceutical research, it is extremely important to perform isolate and identify the protein that is the target of a drug (compound) in vivo. However in the past, isolation and identification of the drug target protein were extremely difficult, and were a problem which required large amounts of time and effort. In order to overcome this problem, through joint research with Professor Hiroshi Handa at the Tokyo Institute of Technology, Tamagawa Seiki has developed the new nanomagnetic particles "FG beads®" and the automated screening system "Target Angler".
FG beads®, are approximately 0.2 μm in diameter and are composed of a plurality ferrite particles coated with a unique polymer called poly-GMA (glycidyl methacrylate). FG beads®, that are manufactured by using this original technology, are used as a carrier for affinity purification and provide characteristics that are superior to conventional carriers, allowing one-step purification of the target proteins. The development of an automated screening system, that magnetically separates and disperses FG beads®, makes it possible to automate the affinity purification process. Concequently, the simultaneous process of multiple samples and time shortening of the process are enabled.
 |
Characteristics of FG beads®
FG beads® are materices for affinity purification that provide superior characteristics which are not available with conventional materices. These characteristics include the following.
- Excellent dispersibility
The nano size of FG beads® gives them excellent dispersibility. The ligands on the surface of the beads efficiently bind to the target protein suspended in solution, yielding a higher recovery rate.
- Large specific surface area
Because of their nano size, FG beads® have a large surface area per unit of volume. This allows many ligands to be immobilized on them, allowing collection of large amounts of the target protein.
- Extremely little non-specific adsorption
The surface of the FG beads® is coated with a special hydrophilic polymer called poly-GMA (glycidyl methacrylate), which results in extremely little non-specific protein adsorption, and allows the target protein to be purified with a high degree of purity.
- Resistant to various organic solvents
FG beads® are resistant to various types of organic solvents, and the beads do not degenerate or dissolve in them. A variety of ligands can be immobilized on the beads in an organic solvent. The organic solvents which can be used include methanol, DMF, DMSO, THF, ethyl acetate, pyridine, dioxane, toluene, dichloromethane, and chloroform.
- Can be magnetically recovered.
Because the FG beads® contain magnetic ferrite, they can be recovered by magnet. Recovery by magnet allows the target proteins to be recovered without effects from factors such as large proteins and degenerated proteins. The use of an automated screening system allows the affinity purification process to be automated, and allows multiple samples to be processed simultaneously, shortening the screening time.
Variety of surface modifications
FG beads® can capture a variety of substances, including chemicals (drugs), proteins, and DNA. From the 13 types of FG beads®, select the type with the optimal surface modification according to the functional group of the substance you want to bind. Because FG beads® are resistant to various organic solvents, they are able to bind a variety of ligands. (Avoid using streptavidin beads or other protein-binding FG beads® in organic solvents.) The beads with ligands immobilized can be used for affinity purification of the target biological substance.
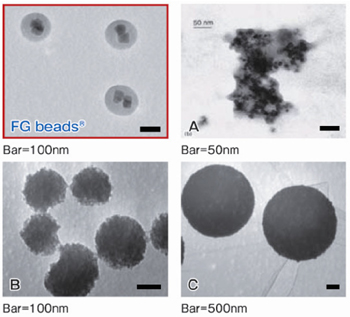
Comparison with other magnetic beads
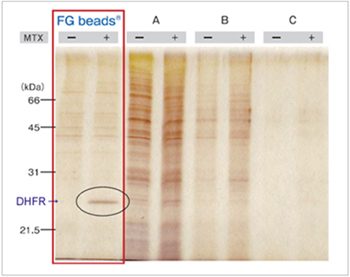
The anti-cancer drug methotrexate (MTX) was immobilized on the magnetic beads of each company using the same method, and affinity purification was performed. Compared with the beads of other companies, FG beads® features less non-specific protein adsorption and allowed purification with a higher recovery rate for the DHFR that was the target protein of MTX.
Affinity purification of MTX binding proteins
Immobilization of MTX on commercial magnetic beads was done in the same manner as in the case of FG beads.

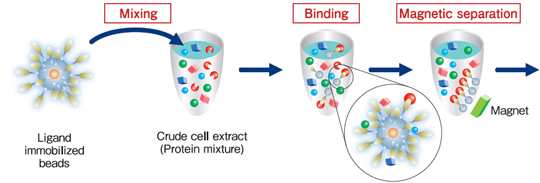
Purification process
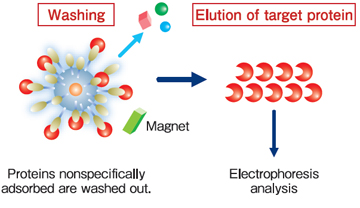
Protein purification (affinity purification) using FG beads® is performed in 3 stages: binding, washing, and elution.
During the binding process, the beads immobilized ligands are mixed with the crude cell extract, and the proteins with affinity for the ligands bind to the beads. During the washing process, the proteins that have bound non-specifically to beads (not the proteins bound specifically to the ligands) are washed off. During the elution process, the specifically-bound proteins are separated from the ligands and recovered. At each of these processes, magnets are used to separate immobilized beads from the crude cell extract, washing buffer, or elution buffer.
Tips; Background Reduction
Immobilization
- Perform centrifugal separation.
The nano size of FG beads gives them excellent dispersibility. As a result, magnetic separation in an organic solvent may be difficult, and it is necessary to recover the FG beads by centrifugal separation.
- Disperse the beads well.
Centrifugal separation causes the FG beads to agglutinate strongly, making it difficult to disperse them. Ordinarily a manual dispersion method or ultrasound would be used to disperse the beads. Although ultrasound separates the beads easily, caution is required due to the possibility of damaging the proteins. Therefore in general it is recommended that ultrasound be used when binding low molecular weight compounds, and that manual dispersion be used when binding proteins. The manual method involves resting the bottom of the micro tube in a plastic test tube stand and moving it roughly to disperse the beads. Depending on the type of micro tube, when using the manual method to disperse the beads, the bottom of the tube may crack or leakage from the lid may occur. Use a micro tube that is strong and has a lid with a tight fit. We recommend the use of cap locks.
Screening
- Perform magnetic separation
When centrifugal separation is performed, heavy proteins and insoluble proteins are also precipitated, raising the level of the background. Because FG beads have high dispersibility, magnetic separation requires time (in some cases 5 minutes or longer). However using magnetic separation avoids the risk of intrusion by these impurities and provides clear results with a low background level.
- Disperse the beads well
If dispersion is insufficient following the FG beads washing process after the binding reaction with the proteins, there is the possibility of impurities remaining inside the bead clusters. Therefore it is necessary to disperse the beads well. Use the manual method to disperse the beads. (With ultrasound, there is the risk that the proteins will be damaged.)
Application Examples
Purification of target protein of Thalidomide (elucidation of the teratogenic mechanism)
CRBN (Cereblon) and DDB1 were isolated from human cell extract using thalidomide fixed beads. As a result, the teratogenic mechanism of thalidomide was elucidated.
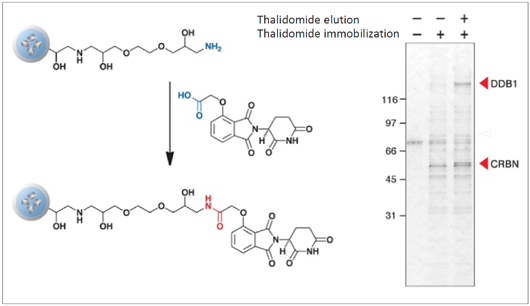
T. Ito et al., Science 327 (2010) 1345
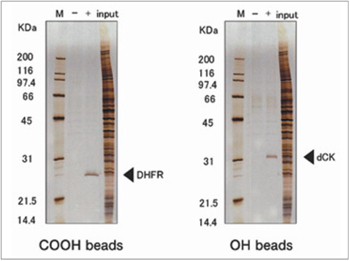
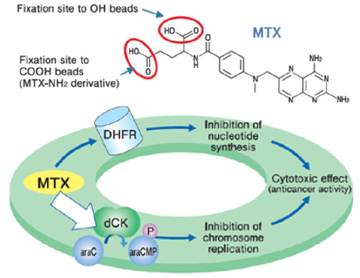
Purification of novel target protein of MTX (methotrexate)
When MTX was fixed via different site, a novel protein was purified and identified as deoxycytidine kinase (dCK). As a result, a possible mechanism of synergistic effect between MTX and ara-C on malignant lymphoma was proposed.
H. Uga et al., Mol. Pharmacol. 70 (2006) 1832
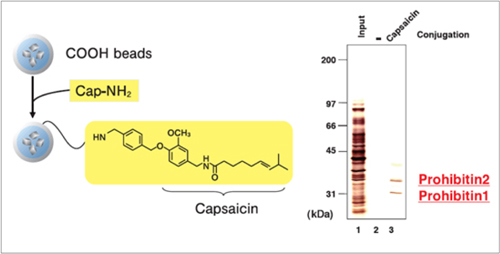
Purification of target proteins of Capsaicin
Prohibitin 1 and prohibitin 2 were isolated from human myeloid leukemia NB4 cell extract using capsaicin derivative (Cap-NH2) fixed beads. As a result, the apoptosis induction mechanism of capsaicin was elucidated.
Elucidation of mechanism of enteropathogenic E. coli enfection
EspB is a protein of enteropathogenic E. coli (EPEC) essential for infection in humans, Myosin was isolated from human cell extract using EspB fixed beads. As a result, the mechanism of EPEC infection was elucidated.
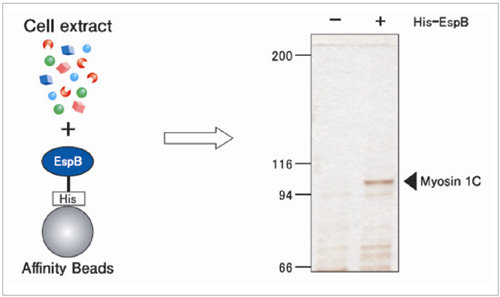
Y. lizumi et al., Cell Host & Microbe. 2 (2007) 383
Application Data: Protein A beads / Protein G beads
Features
-
High recovery IgG binding capacity - more than twice the amount of a competitor.
-
High purity Extremely low non-specific adsorption.
-
Quick processing 30 minutes for IgG binding.
IgG Purification
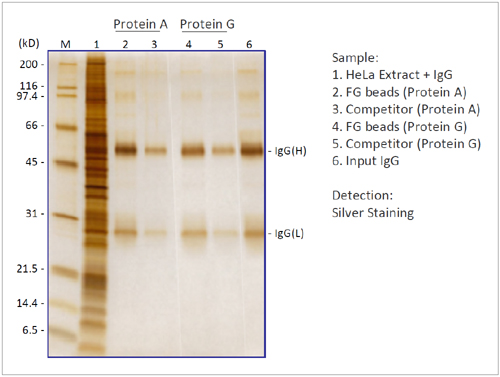
We compared the binding capacity of FG beads to capture IgG in HeLa cell extracts with the beads of a competitor. FG beads captured larger amounts of IgG than the competitor's beads.
- Add 10 µg of IgG into 200 µg of HeLa cell extracts (200 µl).
- Add 0.3 mg of each beads to the HeLa cell extracts.
- React for 10 min at 4°C and separate beads from the HeLa cell extract.
- Elute bound IgG by adding Glycine-HCl.
Immunoprecipitation

We checked the performance of FG beads (Protein A and Protein G) in an immunoprecipitation experiment. By using FG beads, antigen HSA was immunoprecipitated with high recovery and extremely low non-specific adsorption.
- Immobilize anti-Human Serum Albumin antibody on FG beads.
- Add 400 ng of HSA into 200 µg of HeLa cell extracts (200 µl).
- Add 0.1 mg of each beads to HeLa cell extracts.
- React for 120 min at 4°C and separate beads from the HeLa cell extract.
- Elute bound IgG and HSA by adding Glycine-HCl.
Application Data: Streptavidin Beads
Immunoprecipitation
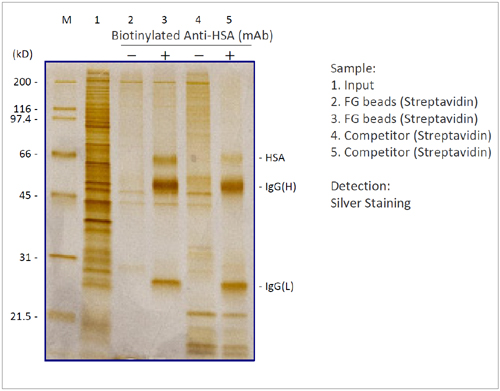
We compared the performance of FG beads in an immunoprecipitation experiment with the beads of a competitor. By using FG beads, antigen HSA was immunoprecipitated with high recovery and extremely low non-specific adsorption.
- Immobolize biotinylated anti-HSA antibody on Streptavidin beads.
- Add 1 µg of HSA into 600 µg of HeLa cell extracts (200 µl).
- Add 0.5 mg of each beads to the HeLa cell extracts.
- React for 60 min at 4°C and separate beads from the HeLa cell extract.
- Elute bound IgG and HSA by adding Glycine-HCl.
Affinity Purification of Drug Target Protein
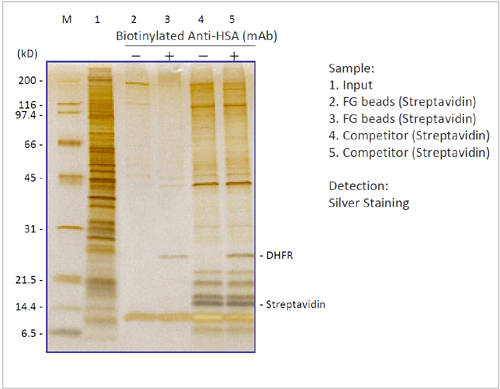
We compared the performance of biotinylated drug MTX (Methotrexate) immbilized FG beads with the beads of a competitor in a target protein purification expertiment. By using FG beads, MTX target protein DHFR was purified with extremely low non-specific adsorption.
- Immobolize biotinylated MTX on Streptavidin beads.
- Add 0.5 mg of each beads into 600 µg of HeLa cell extracts (200 µl).
- React for 120 min at 4°C and separate beads from the HeLa cell extract.
- Elute bound DHFR by adding elution buffer.
Application Data: Azide beads / Alkyne beads
Application; Target fishing
Affinity purification of target protein, DHRF (Dihydrofolate reductase), of MTX (methotrexate) with FG Azide and Alkyne beads.
1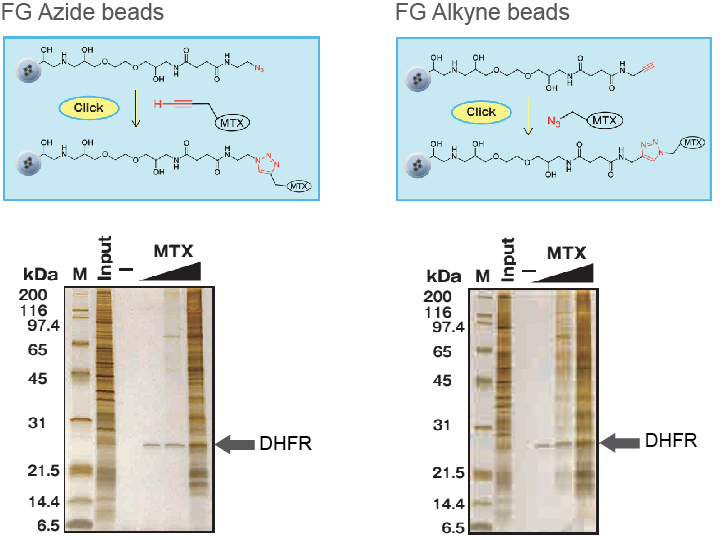
Ordering Information
| Product | Cat.No. | Price | |
|---|---|---|---|
| Please visit "FG beads: Line-up" below | - | - | Inquire |
